Tutorial: water level & salinity data
This tutorial covers water level and salinity monitoring data
We will download the relevant data using this package (python-sa-gwdata) – importable as sa_gwdata – and use some other packages for other things:
matplotlib, numpy, pandas - used in the background
[1]:
import sa_gwdata
import pandas as pd
import matplotlib.pyplot as plt
import geopandas as gpd
# import contextily as cx
[8]:
plt.rcParams['figure.dpi'] = 100
Water level data
I’ll assume that you have already worked out which well you want to download data for - please see previous tutorials for how to find wells. We’re going to look at data from a monitoring well near Port Lincoln: 6028-536 (also known as obswell FLN029).
First we need to make sure we have correctly identified the well, and obtain its drillhole number:
[3]:
wells = sa_gwdata.find_wells("6028-536")
[4]:
wells
[4]:
['FLN029']
[5]:
df = sa_gwdata.water_levels(wells)
df.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 562 entries, 0 to 561
Data columns (total 21 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 dh_no 562 non-null int64
1 network 562 non-null object
2 unit_long 562 non-null int64
3 aquifer 562 non-null object
4 easting 562 non-null float64
5 northing 562 non-null float64
6 zone 562 non-null int64
7 unit_hyphen 562 non-null object
8 obs_no 562 non-null object
9 obs_date 562 non-null datetime64[ns]
10 dtw 561 non-null float64
11 swl 561 non-null float64
12 rswl 561 non-null float64
13 pressure 0 non-null float64
14 temperature 0 non-null float64
15 dry_ind 0 non-null float64
16 anomalous_ind 562 non-null object
17 pump_ind 562 non-null object
18 measured_during 562 non-null object
19 data_source 562 non-null object
20 comments 4 non-null object
dtypes: datetime64[ns](1), float64(8), int64(3), object(9)
memory usage: 92.3+ KB
[6]:
df[['obs_date', 'dtw', 'swl', 'rswl', 'anomalous_ind', 'comments']]
[6]:
| obs_date | dtw | swl | rswl | anomalous_ind | comments | |
|---|---|---|---|---|---|---|
| 0 | 1959-02-26 | 10.67 | 10.67 | 1.67 | N | NaN |
| 1 | 1960-02-11 | 10.76 | 10.76 | 1.58 | N | NaN |
| 2 | 1964-11-04 | 10.70 | 10.70 | 1.64 | N | NaN |
| 3 | 1965-10-06 | 10.96 | 10.96 | 1.38 | N | NaN |
| 4 | 1966-01-06 | 11.01 | 11.01 | 1.33 | N | NaN |
| ... | ... | ... | ... | ... | ... | ... |
| 557 | 2020-10-11 | 12.80 | 12.22 | 0.12 | N | NaN |
| 558 | 2021-03-20 | 12.89 | 12.31 | 0.03 | N | NaN |
| 559 | 2021-10-26 | 12.87 | 12.29 | 0.05 | N | NaN |
| 560 | 2022-04-11 | 12.96 | 12.38 | -0.04 | N | NaN |
| 561 | 2022-10-07 | 12.90 | 12.32 | 0.02 | N | NaN |
562 rows × 6 columns
[10]:
fig = plt.figure()
ax = fig.add_subplot(111)
ax.plot(df.obs_date, df.swl, marker='x')
ax.invert_yaxis()
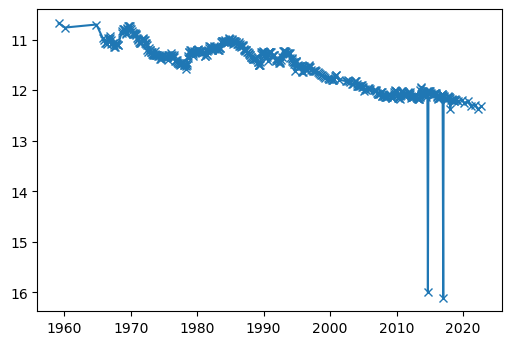
Anomalous data points
Note the three outliers. These have actually been flagged as anomalous:
[11]:
df[df.obs_date.dt.year == 2014][['obs_date', 'dtw', 'swl', 'rswl', 'anomalous_ind', 'comments']]
[11]:
| obs_date | dtw | swl | rswl | anomalous_ind | comments | |
|---|---|---|---|---|---|---|
| 493 | 2014-01-13 | 12.61 | 12.03 | 0.31 | N | NaN |
| 494 | 2014-02-10 | 12.64 | 12.06 | 0.28 | N | NaN |
| 495 | 2014-03-11 | 12.67 | 12.09 | 0.25 | N | NaN |
| 496 | 2014-04-09 | 12.68 | 12.10 | 0.24 | N | NaN |
| 497 | 2014-05-04 | 12.70 | 12.12 | 0.22 | N | NaN |
| 498 | 2014-05-05 | 12.70 | 12.12 | 0.22 | N | NaN |
| 499 | 2014-06-04 | 12.71 | 12.13 | 0.21 | N | NaN |
| 500 | 2014-07-28 | 12.61 | 12.03 | 0.31 | N | NaN |
| 501 | 2014-08-25 | 12.60 | 12.02 | 0.32 | N | NaN |
| 502 | 2014-09-22 | 16.58 | 16.00 | -3.66 | Y | Transcription error |
| 503 | 2014-10-22 | 12.59 | 12.01 | 0.33 | N | NaN |
| 504 | 2014-11-17 | 12.59 | 12.01 | 0.33 | N | NaN |
| 505 | 2014-12-15 | 12.59 | 12.01 | 0.33 | N | NaN |
And if you want, you can opt to always exclude “anomalous” readings, although it is should be noted the definition of “anomalous” is somewhat subjective:
[12]:
df = sa_gwdata.water_levels(wells, anomalous=False)
df[df.obs_date.dt.year == 2014][['obs_date', 'dtw', 'swl', 'rswl', 'anomalous_ind', 'comments']]
[12]:
| obs_date | dtw | swl | rswl | anomalous_ind | comments | |
|---|---|---|---|---|---|---|
| 493 | 2014-01-13 | 12.61 | 12.03 | 0.31 | N | NaN |
| 494 | 2014-02-10 | 12.64 | 12.06 | 0.28 | N | NaN |
| 495 | 2014-03-11 | 12.67 | 12.09 | 0.25 | N | NaN |
| 496 | 2014-04-09 | 12.68 | 12.10 | 0.24 | N | NaN |
| 497 | 2014-05-04 | 12.70 | 12.12 | 0.22 | N | NaN |
| 498 | 2014-05-05 | 12.70 | 12.12 | 0.22 | N | NaN |
| 499 | 2014-06-04 | 12.71 | 12.13 | 0.21 | N | NaN |
| 500 | 2014-07-28 | 12.61 | 12.03 | 0.31 | N | NaN |
| 501 | 2014-08-25 | 12.60 | 12.02 | 0.32 | N | NaN |
| 502 | 2014-10-22 | 12.59 | 12.01 | 0.33 | N | NaN |
| 503 | 2014-11-17 | 12.59 | 12.01 | 0.33 | N | NaN |
| 504 | 2014-12-15 | 12.59 | 12.01 | 0.33 | N | NaN |
Groundwater level parameters
There are three different water level parameters:
Depth to Water (dtw/DTW)
This is in metres measured below a reference point. Increasing numbers indicate an increasing depth to water. Negative numbers indicate flowing artesian conditions. The reference point’s true elevation above ground surface can change i.e. if a casing standpipe is chopped off or installed, so although this number is what is in effect measured in the real world, it should not be used for analysis.
Standing Water Level (swl/SWL)
This is the Depth to Water, automatically corrected such that it represents a depth below ground level. Increasing numbers indicate an increasing depth to water. Negative numbers indicate flowing artesian conditions. The SWL and DTW values are only different when well elevation data is available and the two elevation values (of the reference point, and the ground) are different - the difference is used to calculate the SWL. If they are the same (e.g. maybe the elevation was derived from a digital elevation model), or no elevation data is available, then the SWL will be present but it will be identical to the DTW. I recommend that you always use the SWL value (or RSWL, see below).
Reduced Standing Water Level (rswl/RWL)
This value has been corrected to represent the groundwater level measured above Australian Height Datum (AHD). Increasing numbers indicate a decreasing depth to water. RSWL values are only present when we have an elevation survey for that well. This is obviously the correct value to use for most purposes, if it is available.
Note that there is no distinguishing between physical groundwater levels (i.e. the water table) and water levels as measured in a well which penetrates and provides access to a confined aquifer (i.e. a groundwater pressure level). Even wells which access confined groundwater with flowing artesian conditions, where the pressure level is measured using a pressure gauge at surface, have data reported using this method as “Depth to Water” (which would in that case be a negative number).
To compare all of these parameters, see them on a chart:
[14]:
fig = plt.figure()
ax = fig.add_subplot(111)
ax.plot(df.obs_date, df.dtw, label='dtw', color='tab:blue')
ax.plot(df.obs_date, df.swl, label='swl', color='tab:green', lw=12, alpha=0.3)
ax.plot([], [], label='rswl', color='tab:red', alpha=1, lw=2, ls=':')
ax.invert_yaxis()
span = 2.5
ax_top = 10.6
ax2_top = 1.75
ax.set_ylim(ax_top + span, ax_top)
ax.set_ylabel('DTW & SWL', color='black')
ax2 = ax.twinx()
ax2.plot(df.obs_date, df.rswl, label='rswl', color='tab:red', alpha=1, lw=2, ls=':')
ax2.set_ylim(ax2_top - span, ax2_top)
ax2.set_ylabel('RSWL', color='red')
ax.legend()
fig.savefig("figures/groundwater-level-parameters.png", bbox_inches='tight')
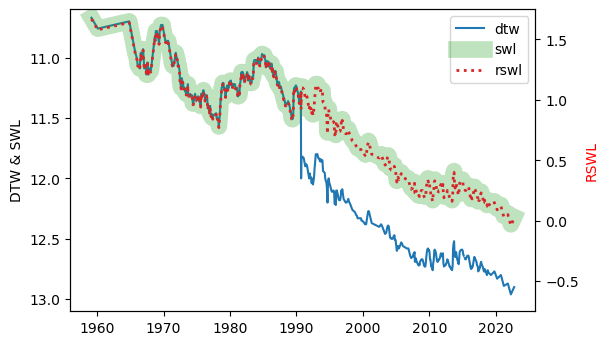
Note the change in casing reference point around 1990 - clearly a casing standpipe was installed, because overnight, the depth to water increased by about one metre, and the new measurements were being made from the top of the standpipe, instead of ground level.
Measured during!
There is a metadata property attached to water level and salinity data called “measured during”. This indicates what kind of activity the data was obtained during, and it is a good way to filter out irrelevant measurements from relevant ones in terms of monitoring.
These are the values that measured_during can take:
[15]:
meas_during_lut = {
'A': 'Aquifer Test',
'D': 'Drilling',
'F': 'Field Survey',
'S': 'Final Sample on drilling completion',
'G': 'Geophysical Logging',
'L': 'Landowner Sample',
'M': 'Monitoring',
'R': 'Rehabilitation',
'U': 'Unknown',
'W': 'Well Yield',
}
An example is from well 6628-5103
[16]:
wells = sa_gwdata.find_wells('6628-5103')
[17]:
df = sa_gwdata.water_levels(wells)
[18]:
df.measured_during.value_counts()
[18]:
M 142
A 3
Name: measured_during, dtype: int64
[19]:
fig = plt.figure()
ax = fig.add_subplot(111)
df[df.measured_during == 'M'].plot(ax=ax, x='obs_date', y='swl', color='tab:blue', marker='+', lw=0.5, label='Monitoring')
df[df.measured_during == 'A'].plot(ax=ax, x='obs_date', y='swl', color='tab:red', marker='x', lw=0.5, label='Aquifer test')
df.plot(ax=ax, x='obs_date', y='swl', lw=0.1, color='grey', label='all data')
ax.legend()
ax.invert_yaxis()
ax.set_ylabel('swl')
[19]:
Text(0, 0.5, 'swl')
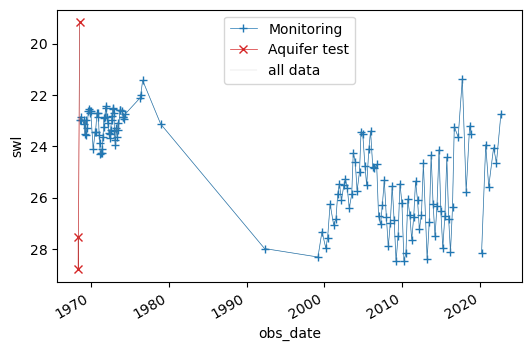
As you can see, filtering to only retain measured during “M” (monitoring) is often a good idea for analysis of water level data.
Salinity data
Salinity tends to be monitored from different wells than water level.
Let’s look at LKW039, a well from the other side of Eyre Peninsula at Coffin Bay.
[20]:
wells = sa_gwdata.find_wells("LKW 39")
[21]:
wells
[21]:
['LKW039']
[22]:
df = sa_gwdata.salinities(wells)
df.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 168 entries, 0 to 167
Data columns (total 20 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 dh_no 168 non-null int64
1 network 168 non-null object
2 aquifer 168 non-null object
3 unit_hyphen 168 non-null object
4 unit_long 168 non-null int64
5 obs_no 168 non-null object
6 collected_date 168 non-null datetime64[ns]
7 collected_time 7 non-null object
8 tds 168 non-null int64
9 ec 168 non-null int64
10 ph 20 non-null float64
11 sample_type 168 non-null object
12 anomalous_ind 168 non-null object
13 test_place 86 non-null object
14 extract_method 165 non-null object
15 measured_during 168 non-null object
16 data_source 168 non-null object
17 easting 168 non-null float64
18 northing 168 non-null float64
19 zone 168 non-null int64
dtypes: datetime64[ns](1), float64(3), int64(5), object(11)
memory usage: 26.4+ KB
These are the important fields:
TDS (Total dissolved solids in mg/L): almost always estimated from the EC measurement by a standard formula - TODO document
EC (Electrical conductivity at 25 deg C in uS/cm): the electrical conductivity, corrected for temperature
Anomalous indicator Y/N: as for water level above
Test place: location that the EC was measured: values include:
U = Unknown
F = During fieldwork
RP = Regency Park i.e. the water test room
Extraction method: how the water sample was obtained for testing.
Measured during: see discussion above
Data source: where the data came from
[23]:
df[['collected_date', 'tds', 'ec', 'anomalous_ind', 'test_place', 'extract_method', 'measured_during', 'data_source']]
[23]:
| collected_date | tds | ec | anomalous_ind | test_place | extract_method | measured_during | data_source | |
|---|---|---|---|---|---|---|---|---|
| 0 | 1985-03-25 | 392 | 712 | N | U | PUMP | W | DEWNR |
| 1 | 1985-03-25 | 392 | 712 | N | U | PUMP | W | DEWNR |
| 2 | 1985-03-25 | 393 | 714 | N | U | PUMP | W | DEWNR |
| 3 | 1985-03-25 | 391 | 710 | N | U | PUMP | W | DEWNR |
| 4 | 1985-03-25 | 397 | 722 | N | U | PUMP | W | DEWNR |
| ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 163 | 2015-04-20 | 1049 | 1900 | N | NaN | PUMP | M | DEWNR |
| 164 | 2016-04-18 | 1077 | 1950 | N | F | PUMP | M | DEWNR |
| 165 | 2019-12-14 | 1127 | 2040 | N | RP | PUMP | M | DEW |
| 166 | 2020-10-18 | 1076 | 1948 | N | RP | PUMP | M | DEW |
| 167 | 2021-10-26 | 1116 | 2020 | N | RP | PUMP | M | DEW |
168 rows × 8 columns
A key field for salinity data is certainly the extraction method, and usually values like PUMP and AIRL (airlifting) are the main ones you should filter for:
[24]:
df.extract_method.fillna("NA").value_counts()
[24]:
BAIL 134
PUMP 31
NA 3
Name: extract_method, dtype: int64
[25]:
fig = plt.figure()
ax = fig.add_subplot(111)
df[df.extract_method == 'PUMP'].plot(
ax=ax, x='collected_date', y='tds', color='tab:blue', marker='s', ms=10, ls='none', label='PUMP', mfc='skyblue'
)
df[df.extract_method == 'BAIL'].plot(
ax=ax, x='collected_date', y='tds', color='tab:orange', marker='^', ms=10, ls='none', label='BAIL', mfc='none'
)
df[pd.isnull(df.extract_method)].plot(
ax=ax, x='collected_date', y='tds', color='black', marker='*', ms=15, ls='none', label='null', mfc='lightgrey'
)
ax.legend()
ax.set_ylabel('tds')
[25]:
Text(0, 0.5, 'tds')
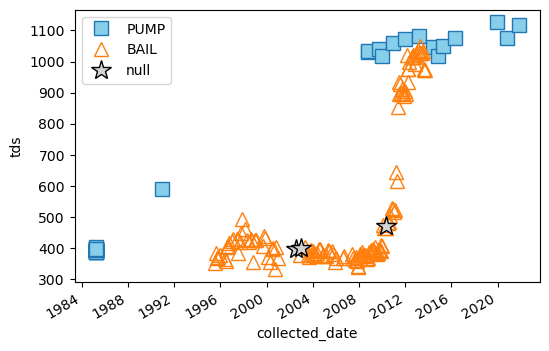
[ ]: